Walkthrough Overview
This walkthrough guides you through the analysis process for a 2022 mobile phone survey conducted in Rwanda. The data for the analysis and the questionnaire metadata can be accessed at the following links and saved on your computer:
The analysis follows the general structure outlined below:
Loading survey data and questionnaire metadata
Labeling and describing data, applying skip logic, and recoding indicators
Analyzing survey data
Exporting results and other outputs
Loading survey data and questionnaire metadata
The survey data can be loaded with
read.csv("ug_publicuse_5421.csv"). Loading the
questionnaire metadata relies on the following three functions:
get_codebook(), get_descriptions(), and
get_skiplogic(). The R code below shows these steps and
their output:
Loading data
rw_publicuse_4483 <- read.csv("rw_publicuse_4483.csv")Importing codebook
The codebook contains each variable (grp) and the
potential response labels (values) and the responses coded
in the data (options).
codebook <- get_codebook(path = "manifest.json")Importing variable descriptions
The descriptions contains a detailed description (desc)
for each variable (var).
descriptions <- get_descriptions(path = "manifest.json")Importing skip logic
A data frame containing the skip logic information, including variables, question IDs, question text, response values, and skip logic conditions.
skip <- get_skiplogic(path = "manifest.json")These inputs are used in data manipulation, analysis, and exporting functions and the above functions are frequently referenced within processing steps.
Labeling and describing data, applying skip logic, and recoding indicators
Label data
The first step in the data manipulation process is to apply data
labels to the survey data using the output from
get_codebook().
rw_label <- label_data(rw_publicuse_4483, codebook = codebook)Applying skip logic
Following the labeling of data, skip logic from the questionnaire is
applied to the survey results to determine when missing values are truly
NA or if they should be recoded as
NOT APPLICABLE. As this function runs, it will output
information in the console that highlights the variable that previous
responses could potentially skip (Skip Logic Variable) and
the parent value and responses that would allow that variable to be
shown to a respondent (Parent variable and
Response values).
rw_label_skip <- apply_skiplogic(path = "manifest.json", x = rw_label)#>
#>
#> ***** Skip Logic Variable: urbanicity *****
#> Parent variable: location
#> Response values: East
#> Response values: West
#> Response values: South
#> Response values: North
#> Response values: Skip
#> Response values: City of Kigali
#>
#>
#> ***** Skip Logic Variable: education *****
#> Parent variable: urbanicity
#> Response values: City
#> Response values: Rural
#> Response values: Skip
#>
#>
#> ***** Skip Logic Variable: smoke1_daily *****
#> Parent variable: smoke1
#> Response values: Yes
#>
#>
#> ***** Skip Logic Variable: smoke1_ever *****
#> Parent variable: smoke1
#> Response values: No
#> Response values: Skip
#>
#>
#> ***** Skip Logic Variable: smoke2 *****
#>
#>
#> ***** Skip Logic Variable: smoke1_everdaily *****
#>
#>
#> ***** Skip Logic Variable: smoke2_ever *****
#> Parent variable: smoke2
#> Response values: No
#> Response values: Skip
#>
#>
#> ***** Skip Logic Variable: ets_home *****
#>
#>
#> ***** Skip Logic Variable: ets_work *****
#> Parent variable: ets_home
#> Response values: Yes
#> Response values: No
#> Response values: Skip
#>
#>
#> ***** Skip Logic Variable: alcohol_freq *****
#> Parent variable: alcohol_12mth
#> Response values: Yes
#>
#>
#> ***** Skip Logic Variable: alcohol1 *****
#> Parent variable: alcohol_freq
#> Response values: Daily
#> Response values: 3-6 days per week
#> Response values: 1-2 days per week
#> Response values: 1-3 days per month
#> Response values: Less than once per month
#> Response values: Skip
#>
#>
#> ***** Skip Logic Variable: vegetable1 *****
#>
#>
#> ***** Skip Logic Variable: fruit2 *****
#> Parent variable: fruit1
#> Response values: from 1 to 7
#> Skip values:
#>
#>
#> ***** Skip Logic Variable: salt1 *****
#>
#>
#> ***** Skip Logic Variable: vegetable2 *****
#> Parent variable: vegetable1
#> Response values: from 1 to 7
#> Skip values:
#>
#>
#> ***** Skip Logic Variable: salt2 *****
#> Parent variable: salt1
#> Response values: Always
#> Response values: Often
#> Response values: Sometimes
#> Response values: Rarely
#> Response values: Never
#> Response values: Skip
#>
#>
#> ***** Skip Logic Variable: salt3 *****
#> Parent variable: salt2
#> Response values: Always
#> Response values: Often
#> Response values: Sometimes
#> Response values: Rarely
#> Response values: Never
#> Response values: Don't Know
#> Response values: Skip
#>
#>
#> ***** Skip Logic Variable: bp1 *****
#> Parent variable: bp_measured
#> Response values: Yes
#>
#>
#> ***** Skip Logic Variable: bp2 *****
#> Parent variable: bp1
#> Response values: YesRecoding variables
Once skip logic is applied, we can then move into recoding variables
or generating new variables to use in the analysis. One example is with
smoke1 and smoke1_ever. smoke1
asks whether a respondent currently smokes tobacco products and
smoke1_ever asks respondents if they have ever smoked
tobacco products. Due to the built-in skip logic, if a person answered
yes to smoke1, they wouldn’t be asked
smoke1_ever because we know that they currently smoke
tobacco products. To get the correct population-level indicator for
smoke1_ever, we need to account for the yes’s
from smoke1. To do this, we can use the
recode_var() function.
rw_analysis <- recode_var(rw_label_skip, vars=c("smoke1", "smoke1_ever"),new_var = smoke1_ever_pop) Running this function will generate a popup window like the image
below. This is where you can copy over the correct values for the new
variable called smoke1_ever_pop.
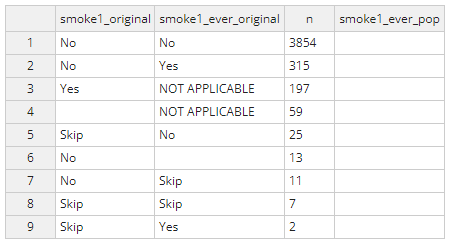
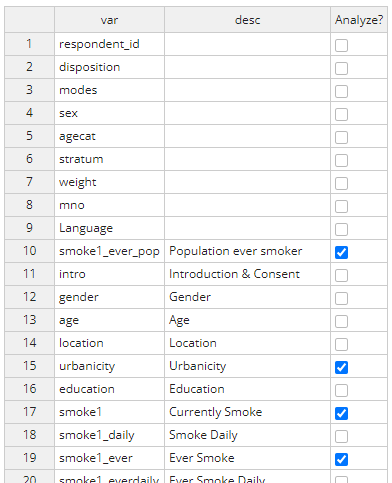
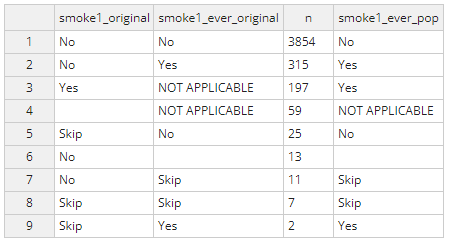
Applying the logic above, the completed table should look like the
table below. Once complete, select ‘done’ at the top right and the
edited dataset will be saved as rw_analysis.
The recode_var() function can be used for any new
variables that need to be created. REMINDER: always write the
output of this function back to the same analysis dataframe to save the
edits.
Setting final descriptions
The final step is to set the final variable descriptions, especially
for any newly developed indicators from the recode_var()
function. The set_descriptions() function will also create
a popup window where you can add or edit descriptions.
desc <- set_descriptions(path="manifest.json", x=rw_analysis)Analyzing survey data
Selecting survey variables for analysis
The first step in this process is to select the variables of interest
for the MPS analysis using the survey_variables() function.
This function will create a popup window like the image below where you
can toggle variables for inclusion in the analysis. Only select the
indicator variables that will correspond to the population-level
estimates and not the variables used for strata/weighting as those are
selected in a different step.
vars <- survey_variables(desc = desc)Creating survey design object
Once the variables are selected, we will use the
mps_design() function to generate the survey design object
using the srvyr package. This function requires the
following information:
df: the analytic dataframeids: the unique identifiers for respondents, usuallyrespondent_idstrata: the strata used for sampling and weighting, typicallyagecatandsexweight: the variable storing the weighting valuevars: the list of variables for the analysis
mps <- mps_design(df = rw_analysis,
ids = respondent_id,
strata = c(agecat, sex),
weight = weight,
vars = vars)The mps object is now a tbl_svy object that
can be used for weighted survey analysis. The object has the following
characteristics:
#> Stratified Independent Sampling design (with replacement)
#> Called via srvyr
#> Sampling variables:
#> - ids: respondent_id
#> - strata: `agecat + sex`
#> - weights: weight
#> Data variables:
#> - respondent_id (chr), agecat (chr), sex (chr), weight (dbl), urbanicity
#> (chr), smoke1 (chr), smoke1_ever_pop (chr)Calculating item non-response rates
When analyzing survey data, item non-response (INR) is an important
factor in evaluating respondent interaction with the survey and
responses for individual questions. The MPSurvey package has a function,
get_inr() that calculates INR for selected variables from
the survey_variables() function. Users can also choose to
include_skips as valid responses by setting this value to
TRUE although most situations should use the default value
of FALSE.
inr <- get_inr(rw_analysis, vars=vars, include_skips = FALSE)The output of this function is a list of two dataframes. The first,
inr, contains one row for each variable of interest with
the percent non-response (perc_non_response), the total
number of valid responses (n_resp), and the total number of
eligible respondents for the question (total). The second
dataframe, responses, shows the total number of responses
for each potential response option for all variables of interest.
Analyze survey
Analyzing the survey can be done using two functions:
analyze_survey() and analyze_survey2(). The
only difference between these is that analyze_survey()
provides overall estimates and estimates by sex where
analyze_survey2() provides overall estimates, estimates by
sex, estimates by agecat, and estimates
stratified by both sex and agecat.
NOTE: to use these functions, the dataframe must have
variables names sex and agecat.
results <- analyze_survey(mps, vars=vars)
results2 <- analyze_survey2(mps, vars=vars)The outputs of these functions are tbl_svysummary
objects that render as formatted tables in the Viewer pane in
RStudio.
results| Characteristic | Overall1 | 95% CI | Female1 | 95% CI | Male1 | 95% CI |
|---|---|---|---|---|---|---|
| urbanicity | ||||||
| City | 2026/4475 45.9% | 44.31%, 47.42% | 973/2151 46.4% | 44.07%, 48.72% | 1053/2324 45.3% | 43.27%, 47.32% |
| Rural | 2449/4475 54.1% | 52.58%, 55.69% | 1178/2151 53.6% | 51.28%, 55.93% | 1271/2324 54.7% | 52.68%, 56.73% |
| smoke1 | 197/4390 4.5% | 3.850%, 5.138% | 71/2106 3.4% | 2.669%, 4.431% | 126/2284 5.6% | 4.688%, 6.584% |
| smoke1_ever_pop | 514/4393 11.7% | 10.69%, 12.69% | 148/2106 7.5% | 6.278%, 8.862% | 366/2287 16.3% | 14.79%, 17.83% |
| agecat | ||||||
| 1 | 1969/4483 38.8% | 38.53%, 39.17% | 992/2155 37.7% | 36.18%, 39.30% | 977/2328 40.1% | 38.73%, 41.46% |
| 2 | 1743/4483 35.1% | 34.81%, 35.38% | 895/2155 34.5% | 33.01%, 36.01% | 848/2328 35.8% | 34.41%, 37.12% |
| 3 | 771/4483 26.1% | 25.46%, 26.67% | 268/2155 27.8% | 25.78%, 29.87% | 503/2328 24.2% | 23.05%, 25.29% |
| 1 n (unweighted)/N (unweighted) % | ||||||
| Abbreviation: CI = Confidence Interval | ||||||
results2| Characteristic |
Overall N = 72508261 |
95% CI |
Female N = 38050421 |
95% CI |
Male N = 34457841 |
95% CI |
1 N = 28168791 |
95% CI |
2 N = 25446241 |
95% CI |
3 N = 18893231 |
95% CI |
Female
|
Male
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
1 N = 14355041 |
95% CI |
2 N = 13125151 |
95% CI |
3 N = 10570231 |
95% CI |
1 N = 13813751 |
95% CI |
2 N = 12321091 |
95% CI |
3 N = 8323001 |
95% CI | |||||||||||||
| urbanicity | ||||||||||||||||||||||||
| City | 2026/4475 45.9% | 44.31%, 47.42% | 973/2151 46.4% | 44.07%, 48.72% | 1053/2324 45.3% | 43.27%, 47.32% | 928/1965 47.2% | 45.02%, 49.43% | 731/1739 42.0% | 39.73%, 44.37% | 367/771 49.0% | 45.13%, 52.85% | 464/990 46.9% | 43.77%, 49.99% | 370/893 41.4% | 38.24%, 44.70% | 139/268 51.9% | 45.88%, 57.80% | 464/975 47.6% | 44.47%, 50.73% | 361/846 42.7% | 39.37%, 46.04% | 228/503 45.3% | 41.02%, 49.71% |
| Rural | 2449/4475 54.1% | 52.58%, 55.69% | 1178/2151 53.6% | 51.28%, 55.93% | 1271/2324 54.7% | 52.68%, 56.73% | 1037/1965 52.8% | 50.57%, 54.98% | 1008/1739 58.0% | 55.63%, 60.27% | 404/771 51.0% | 47.15%, 54.87% | 526/990 53.1% | 50.01%, 56.23% | 523/893 58.6% | 55.30%, 61.76% | 129/268 48.1% | 42.20%, 54.12% | 511/975 52.4% | 49.27%, 55.53% | 485/846 57.3% | 53.96%, 60.63% | 275/503 54.7% | 50.29%, 58.98% |
| smoke1 | 197/4390 4.5% | 3.850%, 5.138% | 71/2106 3.4% | 2.669%, 4.431% | 126/2284 5.6% | 4.688%, 6.584% | 67/1928 3.5% | 2.741%, 4.389% | 89/1708 5.2% | 4.245%, 6.362% | 41/754 4.9% | 3.517%, 6.771% | 31/967 3.2% | 2.263%, 4.524% | 30/874 3.4% | 2.409%, 4.868% | 10/265 3.8% | 2.041%, 6.875% | 36/961 3.7% | 2.713%, 5.151% | 59/834 7.1% | 5.519%, 9.026% | 31/489 6.3% | 4.491%, 8.877% |
| smoke1_ever_pop | 514/4393 11.7% | 10.69%, 12.69% | 148/2106 7.5% | 6.278%, 8.862% | 366/2287 16.3% | 14.79%, 17.83% | 168/1927 8.7% | 7.508%, 10.02% | 208/1713 12.1% | 10.66%, 13.75% | 138/753 15.5% | 13.00%, 18.27% | 56/969 5.8% | 4.473%, 7.438% | 67/874 7.7% | 6.077%, 9.628% | 25/263 9.5% | 6.501%, 13.70% | 112/958 11.7% | 9.804%, 13.89% | 141/839 16.8% | 14.42%, 19.49% | 113/490 23.1% | 19.54%, 27.00% |
| agecat | ||||||||||||||||||||||||
| 1 | 1969/4483 38.8% | 38.53%, 39.17% | 992/2155 37.7% | 36.18%, 39.30% | 977/2328 40.1% | 38.73%, 41.46% | ||||||||||||||||||
| 2 | 1743/4483 35.1% | 34.81%, 35.38% | 895/2155 34.5% | 33.01%, 36.01% | 848/2328 35.8% | 34.41%, 37.12% | ||||||||||||||||||
| 3 | 771/4483 26.1% | 25.46%, 26.67% | 268/2155 27.8% | 25.78%, 29.87% | 503/2328 24.2% | 23.05%, 25.29% | ||||||||||||||||||
| sex | ||||||||||||||||||||||||
| Female | 992/1969 51.0% | 48.75%, 53.17% | 895/1743 51.6% | 49.23%, 53.92% | 268/771 55.9% | 52.26%, 59.57% | 992/992 100.0% | 100.00%, 100.00% | 895/895 100.0% | 100.00%, 100.00% | 268/268 100.0% | 100.00%, 100.00% | ||||||||||||
| Male | 977/1969 49.0% | 46.83%, 51.25% | 848/1743 48.4% | 46.08%, 50.77% | 503/771 44.1% | 40.43%, 47.74% | 977/977 100.0% | 100.00%, 100.00% | 848/848 100.0% | 100.00%, 100.00% | 503/503 100.0% | 100.00%, 100.00% | ||||||||||||
| strat | ||||||||||||||||||||||||
| Female | 992/992 100.0% | 100.00%, 100.00% | 895/895 100.0% | 100.00%, 100.00% | 268/268 100.0% | 100.00%, 100.00% | ||||||||||||||||||
| Male | 977/977 100.0% | 100.00%, 100.00% | 848/848 100.0% | 100.00%, 100.00% | 503/503 100.0% | 100.00%, 100.00% | ||||||||||||||||||
| 1 n (unweighted)/N (unweighted) % | ||||||||||||||||||||||||
| Abbreviation: CI = Confidence Interval | ||||||||||||||||||||||||
Exporting results and other outputs
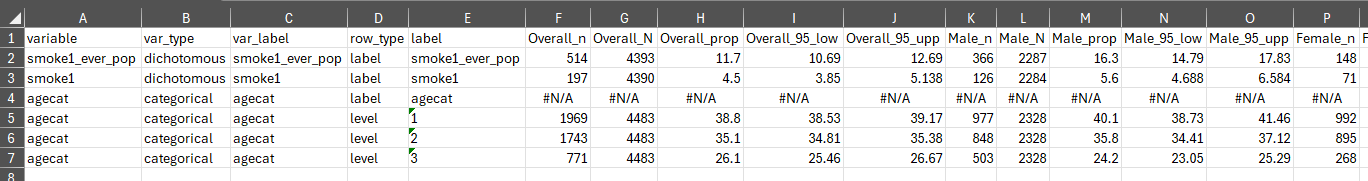
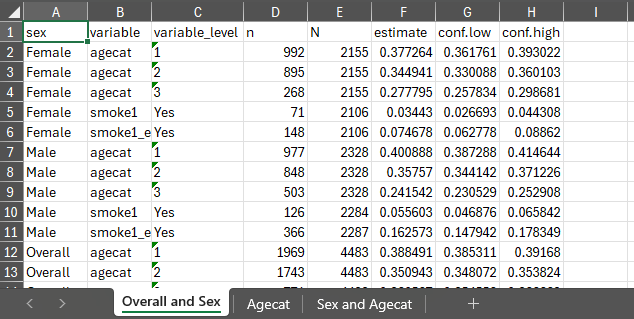
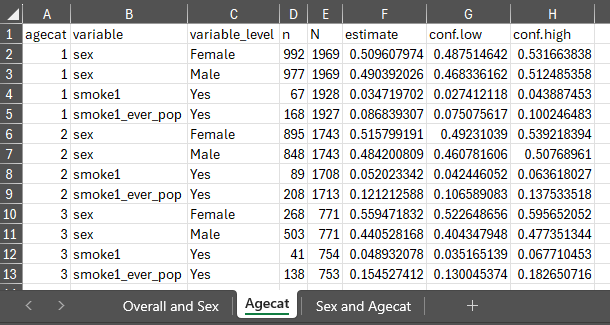
Exporting results
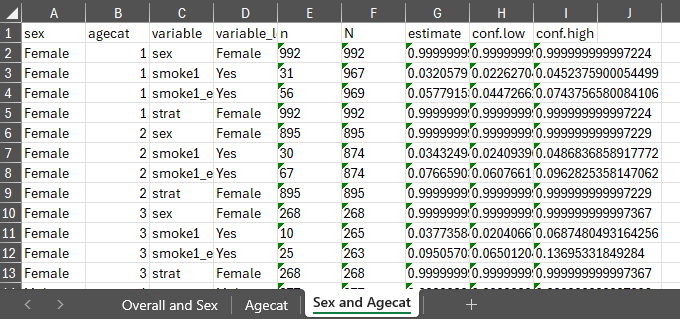
With the previously generated results stored in results
and results2 we can export the generated estimates to excel
using export_results() and export_results2()
in the respective stored tbl_svysummary objects.
export_results(results = results, file = "rw_results.xlsx")
export_results2(results = results2, file = "rw_results2.xlsx")Visualizing questionnaire skip logic
The final function in the MPSurvey package is
visualize_quex(). This function uses the questionnaire skip
logic from the JSON file to create a directed graph showing the flow of
questions and their skip patterns. You can also optionally include a
value for responses to show the number of responses that
flow in each pathway. This value can be directly used from the output of
the get_inr() function by using
inr$responses.
visualize_quex(path = "manifest.json",
file = "rw_quex.pdf",
responses = TRUE,
analysis = rw_analysis)